Tissue-of-Origin Classifier
Single-sample tissue-of-origin inference using an AutoGluon ensemble trained on AACR GENIE v18 data.
Overview
Predict the tissue of origin for a tumor sample using somatic mutations, copy-number alterations, structural variants, and clinical features. The classifier uses a CatBoost + XGBoost ensemble via AutoGluon, integrating 7 feature modalities into ~4,320 features across 22 tumor types, with rank-based CNA normalization for cross-platform compatibility.
Output is a self-contained HTML report with top-3 predictions, full probability distributions, SHAP explanations, and a summary of the input data.
Performance
Evaluation
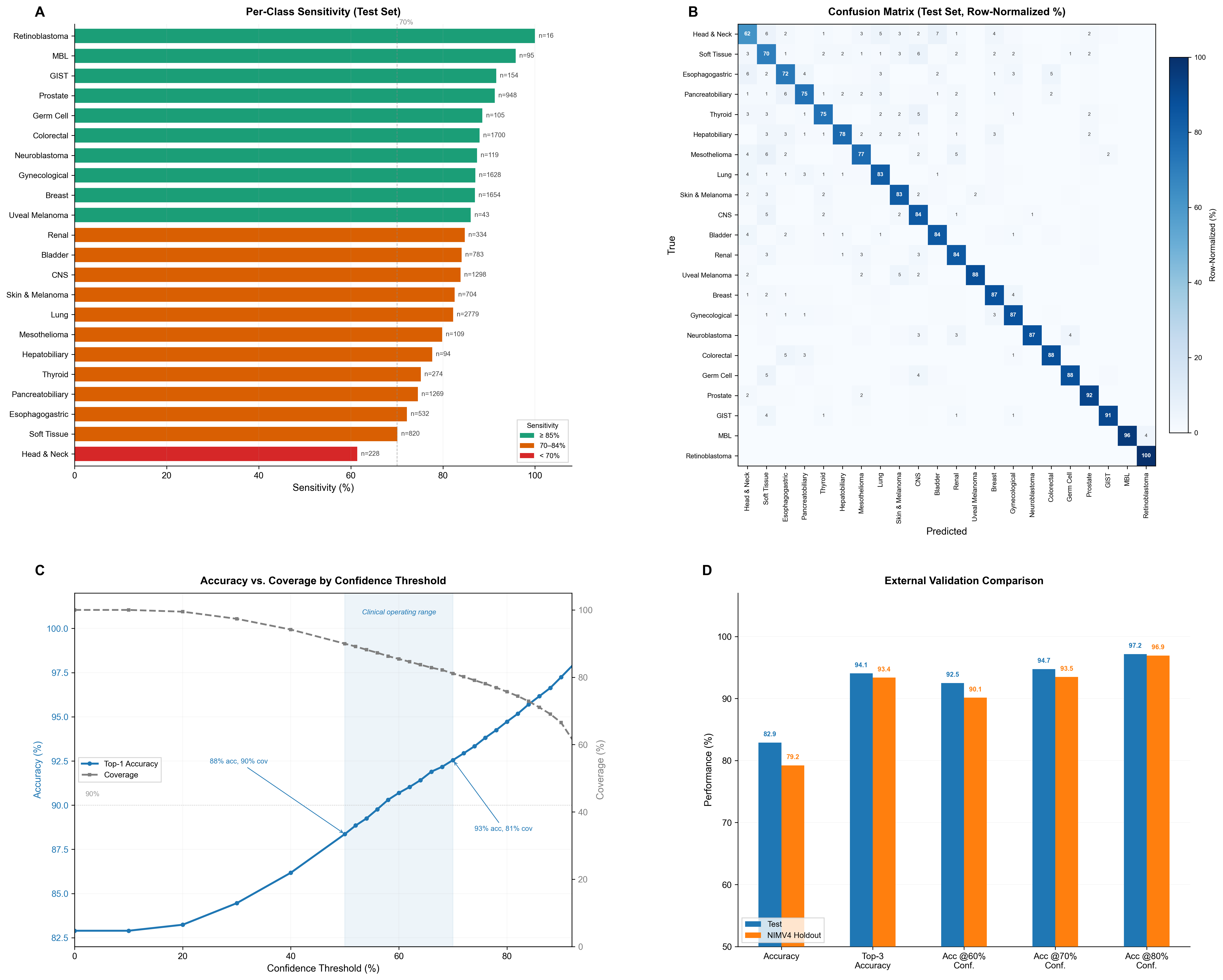
Feature modalities
- Somatic mutations (gene-level)
- Copy-number alterations (arm & gene-level)
- Structural variants
- SBS-6 mutation spectrum
- Mutation frequency
- TERT promoter status
- Clinical features (age, sex)
Installation
Requires Python ≥ 3.11 and uv.
git clone https://github.com/viktorlj/tissue-classifier.git
cd tissue-classifier
uv venv && uv syncUsage
# Basic prediction
tissue-classifier predict \
--maf sample.maf \
--seg sample.seg \
--age 65 --sex Male
# With all options
tissue-classifier predict \
--maf sample.maf \
--seg sample.seg \
--age 65 --sex Male \
--genome hg19 \
--output ./results \
--sample-id PATIENT_001
# Validate input files
tissue-classifier validate --maf sample.maf --seg sample.seg
# Show model info
tissue-classifier infoInput files
MAF (required)
Tab-delimited mutation annotation file with columns:
Hugo_Symbol, Chromosome,
Start_Position, End_Position,
Reference_Allele, Tumor_Seq_Allele2,
Variant_Classification, Variant_Type,
Tumor_Sample_Barcode.
SEG (optional)
Copy-number segmentation file with columns:
ID, chrom, loc.start,
loc.end, seg.mean.
SV (optional)
Structural variant file with columns:
Sample_Id, Site1_Hugo_Symbol,
Site2_Hugo_Symbol.